|
The composite interaction network used for the prediction can also be exported in standard formats, e.g. The plugin can produce a prediction report that lists the details of the query list, source networks and the predicted genes. Users can build their query list using the auto-completion feature, which finds genes by prefix as the user types or by pasting in large gene lists from other sources, such as text files. Users can supply a mixture of symbols from different sources and the plugin will attempt to map them to the corresponding gene. The plugin recognizes gene identifiers, symbols and non-ambiguous synonyms from Entrez, Ensembl, RefSeq, TAIR and Uniprot. However, once the database is downloaded, it does not need to be downloaded again, unless there is an update. all data for human is currently 1.4 GB compressed). 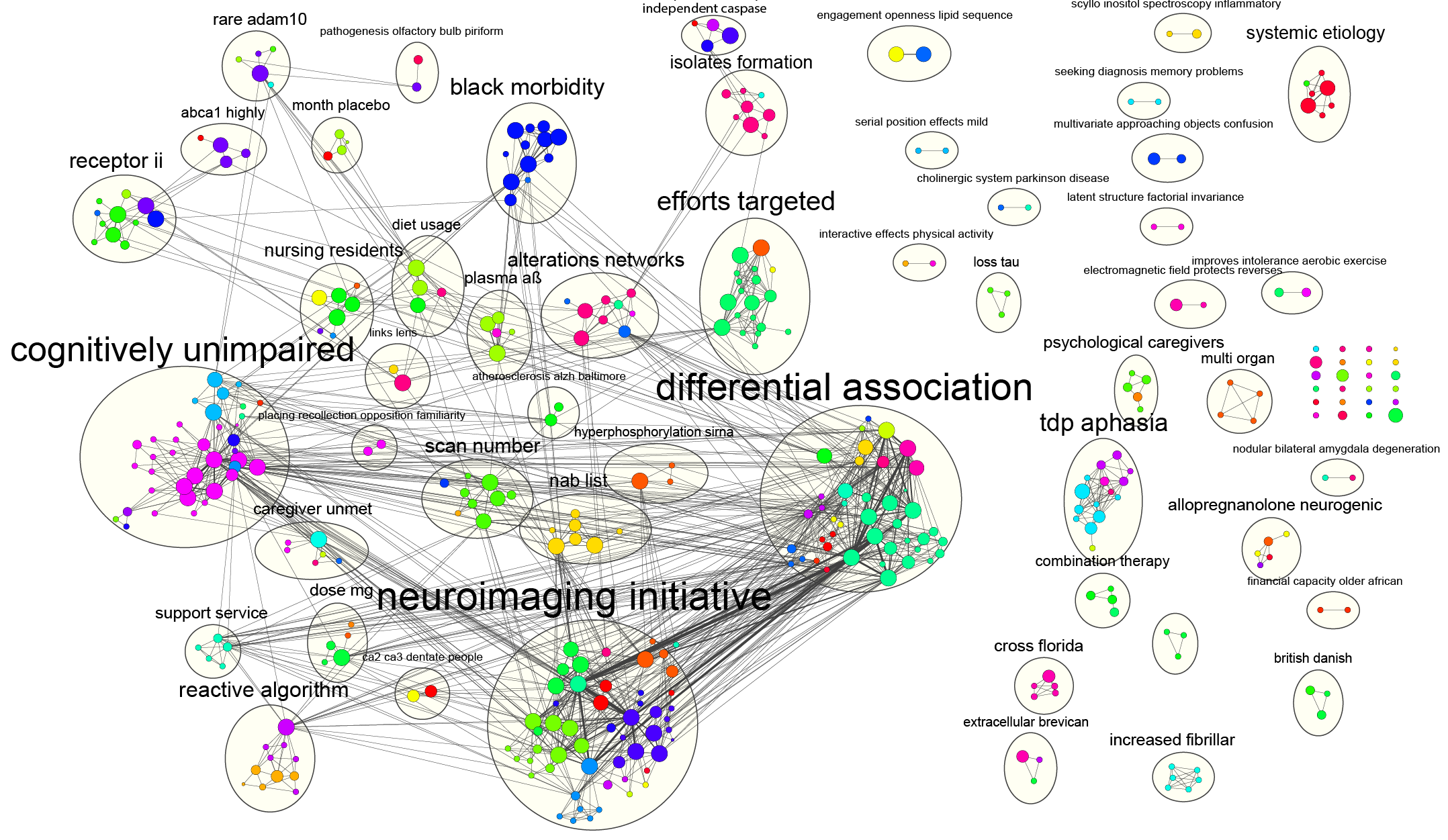
A simple graphical interface aids in this process, which can be time consuming depending on organism choice (e.g. Upon first use, a user must download the latest version of GeneMANIA data for the organisms they are interested in. It can be installed using Cytoscape's menu-driven plugin manager. The plugin automatically checks for these updates and prompts the user to download and install new networks and organisms as they become available.Įase of Use: the GeneMANIA Cytoscape plugin has a user-friendly graphical user interface, which makes powerful prediction tools and data accessible to typical biologists. The data come from a wide range of sources including individual studies and large databases such as BIOGRID (Breitkreutz et al., 2008), GEO (Barrett et al., 2009), I2D (Brown and Jurisica, 2005) and Pathway Commons ( ). The networks are grouped into six categories: co-expression, co-localization, genetic interaction, physical interaction, predicted and shared protein domain. 
The plugin uses a large dataset of functional association networks, which includes over 800 networks for six organisms: Arabidopsis thaliana, Caenorhabditis elegans, Drosophila melanogaster, Mus musculus, Homo sapiens and Saccharomyces cerevisiae. A set of known DNA repair genes were provided as a query (gray nodes) and a number of additional DNA repair genes were predicted to be related (white nodes). The GeneMANIA Cytoscape plugin analysis view showing an example prediction. The resulting predicted network of functional relationships among query and predicted genes is then available as an annotated Cytoscape network for further analysis ( Fig. The plugin extends the Cytoscape network visualization and analysis platform (Shannon et al., 2003) and the functionality of the GeneMANIA gene function prediction website (Warde-Farley et al., 2010) to enable computational biologists and biologists to conduct queries using any number of genes and networks as long as their machine has enough memory. GeneMANIA has been shown to be as good or better in speed and accuracy compared with other gene function prediction algorithms in a competition based on mouse functional association network data (Pena-Castillo et al., 2008). The plugin implements the GeneMANIA algorithm (Mostafavi et al., 2008), which uses a guilt-by-association approach to derive predictions from a combination of potentially heterogeneous data sources. 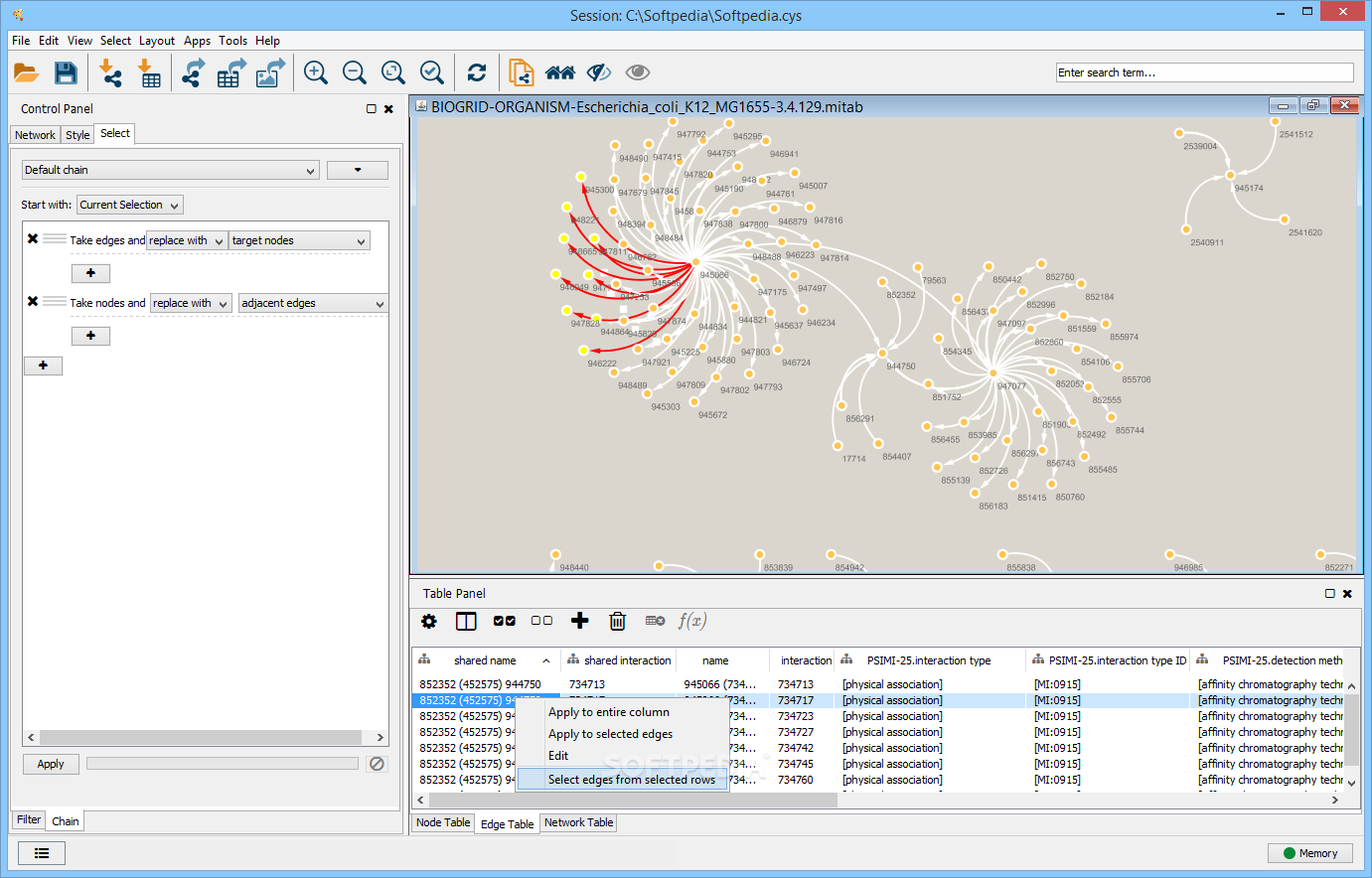
The GeneMANIA Cytoscape plugin is a standalone tool for making fast and efficient gene function predictions.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
